For example, RFLP analysis showed that genetic diversity for SI species far exceeds that of SC species, estimated at 75% versus 7% (Miller and Tanksley, 1990). Species boundaries and genetic diversity have been extensively studied in tomato using a wide range of molecular data (Peralta et al., 2008 and Grandillo et al., 2011). Especially at the geographic margins of the distributions, inter-species changes in incompatibility systems that promote inbreeding over out-crossing have been documented (Peralta et al., 2008 Grandillo et al., 2011). 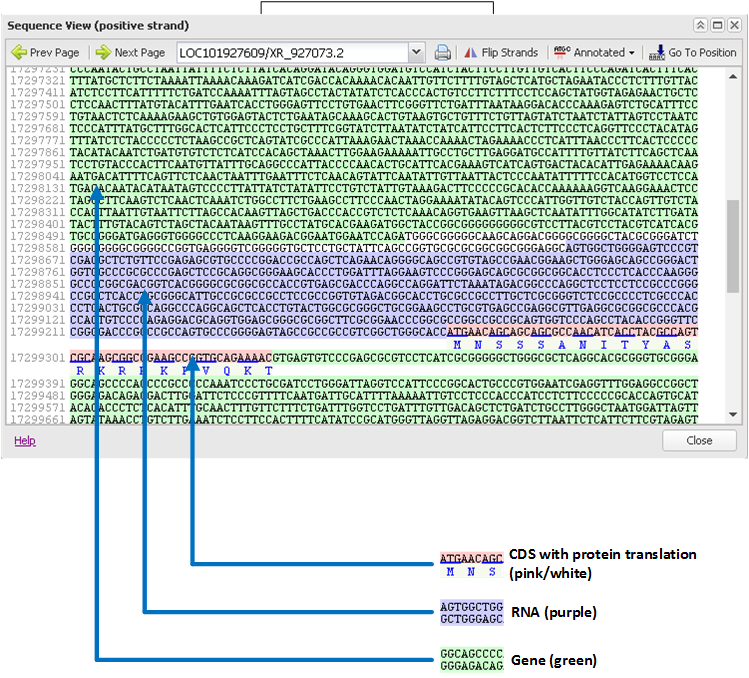
Introgression breeding is possible as cultivated tomato and related wild species are intra-crossable, and most of wild species are also inter-crossable (Rick, 1979, 1986 Spooner et al., 2005) despite the fact that diverse mating systems have evolved, varying from allogamous self-incompatible (SI) and facultative allogamous, to autogamous self-compatible (SC). The first step of introgressive hybridization involves crosses of the cultivated tomato with heirloom species, wild relatives or more distant species of the tomato clade. To enlarge the genetic basis, breeders now focus on introgression of desirable genes from wild relatives into the elite cultivars, but so far, this has been quite limited (Bai and Lindhout, 2007 Singh, 2007). However, the reduced genetic variation that resulted from extensive inbreeding has decelerated tomato crop improvement. These challenges have pushed tomato breeding efforts towards better biotic and abiotic stress tolerance, higher productivity, and increased sensory and nutritional value. The relative small genetic variation became apparent in the face of rapidly changing environmental conditions, competing claims for arable lands, and new consumer requests. In more recent times, tomato has been adapted to different growing systems by adjustment of a small number of traits, including self-pruning, plant height, earliness, fruit morphology and fruit color (Rodríguez et al., 2011 Bauchet and Causse, 2012). As a result, its genetic basis has been seriously narrowed, known as the ‘domestication syndrome’ (Hammer, 1985 Doebley et al., 2006 Bai and Lindhout, 2007 Bauchet and Causse, 2012). However, domestication of tomato is clearly distinct from the species divergence by natural selection as a consequence of selecting for a limited set of traits, including fruit shape and size (Rodríguez et al., 2011). Its economic success is reflected by the fact that, on a global scale, tomato is one of the most important vegetable crops, with a worldwide production of 161 million tonnes covering some 4 800 000 ha ( ). Tomato breeding over recent decades has focused on higher productivity and adaption to different cultivation systems. Solanum is the largest genus in the family, and includes tomato ( Solanum lycopersicum) and various other species of economic importance. Its species occur worldwide, and range from large forest trees in wet rain forests to annual herbs in deserts (Knapp, 2002). The Solanaceae or nightshade family consists of more than 3000 species with great diversity in terms of habit, habitat and morphology. Finally, results suggest that phylogenetic relationships are correlated with habitat, indicating the occurrence of geographical races within these groups, which is of practical importance for Solanum genome evolution studies. 
Using whole-genome SNP information for maximum-likelihood analysis, we achieved complete tree resolution, whereas maximum-likelihood trees based on SNPs from ten fruit and growth genes show incomplete resolution for the crop accessions, partly due to the effect of heterozygous SNPs. In addition, the highest levels of heterozygosity were found for allogamous self-incompatible wild species, while facultative and autogamous self-compatible species display a lower heterozygosity level. 20-fold higher than found in most of the crop accessions, indicating dramatic genetic erosion of crop and heirloom tomatoes. In wild species, the number of single-nucleotide polymorphisms (SNPs) exceeds 10 million, i.e. Using gene models from the annotated Heinz 1706 reference genome, we observed differences in the ratio between non-synonymous and synonymous SNPs (dN/dS) in fruit diversification and plant growth genes compared to a random set of genes, indicating positive selection and differences in selection pressure between crop accessions and wild species. Comparative sequence alignment revealed group-, species- and accession-specific polymorphisms, explaining characteristic fruit traits and growth habits in the various cultivars. Three new reference genomes were reconstructed to support our comparative genome analyses. We explored genetic variation by sequencing a selection of 84 tomato accessions and related wild species representative of the Lycopersicon, Arcanum, Eriopersicon and Neolycopersicon groups, which has yielded a huge amount of precious data on sequence diversity in the tomato clade.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
